Hello,
I am hoping to model a chemical reaction network within a cell, where an organism may change the rate of a reaction given an external stimulus at time t. Is it possible to model a CRN wherea reaction constant, specified in rates changes at time t? Code pasted below.
using Catalyst
using Plots
using DifferentialEquations
rn = @reaction_network begin
a, K --> RP + K
b, RP --> P + RP
(c, d), P + K <--> KP
e, P --> 0
f, RP --> 0
end a b c d e f
rates = (
:a => 0.1,
:b => 0.001,
:c => 0.01,
:d => 0.1,
:e => 0.1,
:f => 0.1
)
tspan = (0., 1000.)
u0 = [
:K => 10,
:RP => 0,
:P => 0,
:KP => 0
]
oprob = ODEProblem(rn, u0, tspan, rates)
solution = solve(oprob, Tsit5(), saveat=10.)
plot(solution)
Concretely, is it possible to generate a piecewise-like function in which the rate constant e changes values at a given time t?
Thank you!
Graeme
If there is a single change of rate at a know moment you could do something like this, where you change the value of a at t=1000 to 1.0
But probably there are more elegant solutions.
rates = (
:a => 0.1,
:b => 0.001,
:c => 0.01,
:d => 0.1,
:e => 0.1,
:f => 0.1
)
tspan = (0., 1000.)
u0 = [
:K => 10,
:RP => 0,
:P => 0,
:KP => 0
]
oprob = ODEProblem(rn, u0, tspan, rates)
solution = solve(oprob, Tsit5(), saveat=10.)
rates = (
:a => 1,
:b => 0.001,
:c => 0.01,
:d => 0.1,
:e => 0.1,
:f => 0.1
)
tspan = (1000., 2000.)
u0 = [
:K => solution.u[end][1],
:RP => solution.u[end][2],
:P => solution.u[end][3],
:KP => solution.u[end][4]
]
oprob = ODEProblem(rn, u0, tspan, rates)
solutionpart2 = solve(oprob, Tsit5(), saveat=10.)
and then combine the two solution parts
There are a bunch of ways to achieve this. Catalyst doesn’t support events yet, but you could explicitly convert rn to a ModelingToolkit ODESystem and pass in a symbolic event. You could also just add an event via the DifferentialEquations.jl interface, see here.
Finally, you could use a simple function for the rate expression to manually flip the rate at a specified time (note I then also set tstops in the call to solve to ensure the ODE solver steps to the switching time exactly):
using Catalyst, OrdinaryDiffEq, Plots
rate(k1,k2,tswitch,t) = (t < tswitch) ? k1 : k2
# this ensures our custom function works with ModelingToolkit/Symbolics.jl
@register rate(k1,k2,tswitch,t)
rn = @reaction_network begin
rate(k1,k2,tswitch,t), A --> 0
end k1 k2 tswitch
p = (:k1 => 0.0, :k2 => 1.0, :tswitch => 1.0)
u0 = [:A => 10.0]
tspan = (0.0,3.0)
oprob = ODEProblem(rn,u0,tspan,p)
sol = solve(oprob, Tsit5(), tstops=[1.0])
plot(sol)
giving
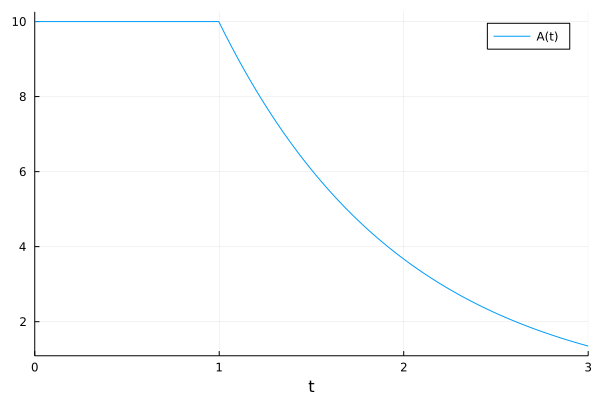
And here is a version that uses IfElse.ifelse, and so probably works better in applications that might need AD:
using Catalyst, OrdinaryDiffEq, Plots, IfElse
rate(k1,k2,tswitch,t) = IfElse.ifelse(t < tswitch, k1, k2)
rn = @reaction_network begin
rate(k1,k2,tswitch,t), A --> 0
end k1 k2 tswitch
p = (:k1 => 0.0, :k2 => 1.0, :tswitch => 1.0)
u0 = [:A => 10.0]
tspan = (0.0,3.0)
oprob = ODEProblem(rn,u0,tspan,p)
sol = solve(oprob, Tsit5(), tstops=[1.0])
plot(sol)
Fantastic. I ended up using IfElse, and it did what I needed. Thank you!
